East Coast Fever (ECF), caused by Theileria parva, is a major constraint to improved livestock keeping in east and central Africa, including Zambia. To understand the dynamics and determine the candidates for immunization in Zambia’s Chongwe and Chisamba districts, a combination of Tp1 and Tp2 gene sequencing and microsatellite analysis using nine markers was conducted from which an abundance of Muguga, Kiambu, Serengeti and Katete epitopes in the field samples was obtained. Phylogenetic analysis showed six (Tp1) and three (Tp2) clusters with an absence of geographical origin clustering. The majority of haplotypes were related to Muguga, Kiambu, Serengeti and Katete, and only a few were related to Chitongo. Both antigens showed purifying selection with an absence of positive selection sites. Furthermore, low to moderate genetic differentiation was observed among and within the populations, and when vaccine stocks were compared with field samples, Chongwe samples showed more similarity to Katete and less to Chitongo, while Chisamba samples showed similarity to both Katete and Chitongo and not to Muguga, Kiambu or Serengeti. We conclude that the use of Katete stock for immunization trials in both Chongwe and Chisamba districts might produce desirable protection against ECF.
Sequence Diversity of Tp1 and Tp2 Antigens and Population Genetic Analysis of Theileria parva in Unvaccinated Cattle in Zambia’s Chongwe and Chisamba Districts
Categories:
Related Post
PREQUALIFICATION OF SUPPLIERS TO SUPPLY VARIOUS PRODUCTS AND SERVICES FOR THE FINANCIAL YEAR 2021 TO THE CENTRE FOR TICKS AND TICK-BORNE DISEASESPREQUALIFICATION OF SUPPLIERS TO SUPPLY VARIOUS PRODUCTS AND SERVICES FOR THE FINANCIAL YEAR 2021 TO THE CENTRE FOR TICKS AND TICK-BORNE DISEASES
The Centre for Ticks and Tick-Borne Diseases (CTTBD) was established by the African Union as a self-financingTrust to assist with the management of ticks and tick-borne diseases on the African

CTTBD TRAINS ECF VACCINATORS IN BURUNDICTTBD TRAINS ECF VACCINATORS IN BURUNDI
One of the core mandates of CTTBD is to conduct tailored courses and training in Animal health, diagnostics and tick-borne disease control. In mid-2019, CTTBD entered into a partnership with
Support of Theileria vaccine production at the Centre for Ticks and Tick Borne Diseases (CTTBD, Malawi) and development of an in vitro potency method [CTTBD-GALVmed]Support of Theileria vaccine production at the Centre for Ticks and Tick Borne Diseases (CTTBD, Malawi) and development of an in vitro potency method [CTTBD-GALVmed]
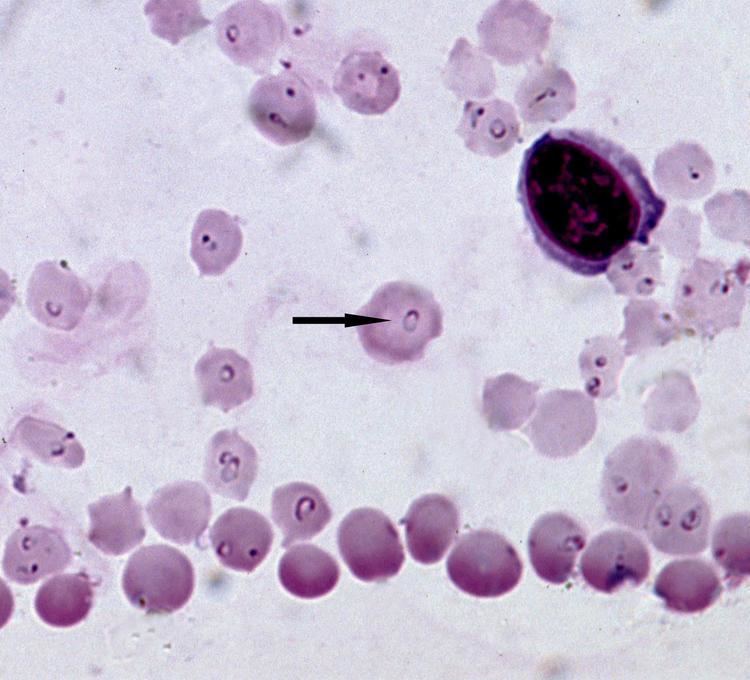
In short Theileria parva is a parasite which kills over 1 million heads of livestock annually in sub-Saharan Africa. Rhipicephalus appendiculatus, the brown ear tick, is the vector of this
